Project 2: Local Feature Matching
0.Project Description
The purpose of this project is to compute the keypoints matching of two pictures. Tha algorithm includes several steps: first, detect basic interests points of the two images by applying Harris corner detector. Second, get the SIFT descriptors of two images according to their corresponding interests points. Third, matching the two features using the nearest neighbor distance ratio to find out those points above certain threshold.
- Get interests points.
- Get features.
- Features matching.
- Other examples.
- Reflection.
1.Get Interests points
Applying Harris Corner Detection in the following algorithm:
- Compute the horizontal and vertical derivates of image Ix and Iy by convolving them with derivatives of Gaussians.
- Compute Ix * Ix, Iy * Iy and Ix * Iy.
- Convolve the three products with a larger Gaussian.
- Compute the Harris value by the following equation.
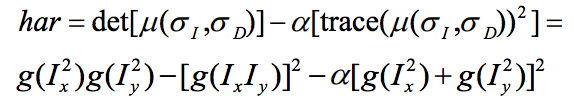
Harris = Ix2 .* Iy2 - Ixy .^ 2 - k .* (Ix2 + Iy2) .* (Ix2 + Iy2);Also, in order to prevent index out of bound because of offset from the interests points to its surrounding, I add a border mask to the Harris matrix:
borderMask = zeros(imSize); borderMask(1 + feature_width:end - feature_width, 1 + feature_width:end - feature_width) = 1; Harris = Harris .* borderMask; - Find local maxima above a certain threshold and store them as detected interests points.
Threashold = 0.0001; k = 0.06; threasholdMask = Harris > Threashold; connected = bwconncomp(threasholdMask); num = connected.NumObjects; pixelIdxList = connected.PixelIdxList; x = zeros(num, 1); y = zeros(num, 1); for i = 1:num currentInd = pixelIdxList{i}; currentPix = Harris(currentInd); [localmax, maxID] = max(currentPix); [j,k] = ind2sub(imSize, currentInd(maxID)); x(i) = k; y(i) = j; end
Gaussian = fspecial('gaussian', 3, 0.1);
[Gx, Gy] = gradient(Gaussian);
Ix = imfilter(image, Gx);
Iy = imfilter(image, Gy);
LargerGaussian = fspecial('gaussian', 9, 1);
Ix2 = imfilter(Ix.*Ix, LargerGaussian);
Iy2 = imfilter(Iy.*Iy, LargerGaussian);
Ixy = imfilter(Ix.*Iy, LargerGaussian);
2.Get features
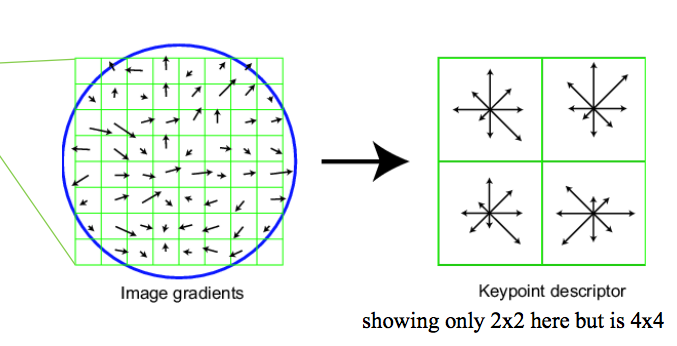
In this step, the local descriptors are returned for those interests points by applying Scale invariant feature transform(SIFT). In my implementation, I construct 4x4 grid cells around the interest points. Then, compute the normalized orientation in 8 directions. Therefore, total of 4 x 4 x 8 = 128 dimensions of descriptors were returned.
features = zeros(length(x), 128);
gaussian = fspecial('gaussian', feature_width, 1);
[Gx, Gy] = gradient(gaussian);
cellSize = feature_width / 4;
Ix = imfilter(image, Gx);
Iy = imfilter(image, Gy);
% calculate orientation of pixels
orientation = atan2(-Iy,Ix)*180/pi;
magnitute = sqrt(Ix .* Ix + Iy .* Iy);
reduce_g = fspecial('gaussian', feature_width, feature_width/2);
for i = 1:length(x)
currentX = x(i) - 2*cellSize;
for j = 1:4
currentY = y(i) - 2*cellSize;
for s = 1:4
currentCell = orientation(currentY:(currentY + cellSize - 1), currentX:(currentX + cellSize - 1));
currentMag = magnitute(currentY:(currentY + cellSize - 1), currentX:(currentX + cellSize - 1));
currentMag = imfilter(currentMag, reduce_g);
for k = -4:1:3
mask = currentCell >= k*45 & currentCell < (k+1)*45;
mag_masked = currentMag(mask);
histo = nnz(mag_masked);
features(i, (j - 1) * 32 + (s - 1) * 8 + k + 5) = histo;
end
currentY = currentY + cellSize;
end
currentX = currentX + cellSize;
end
total = sum(features(i,:));
features(i, :) = features(i, :)/total;
end
end
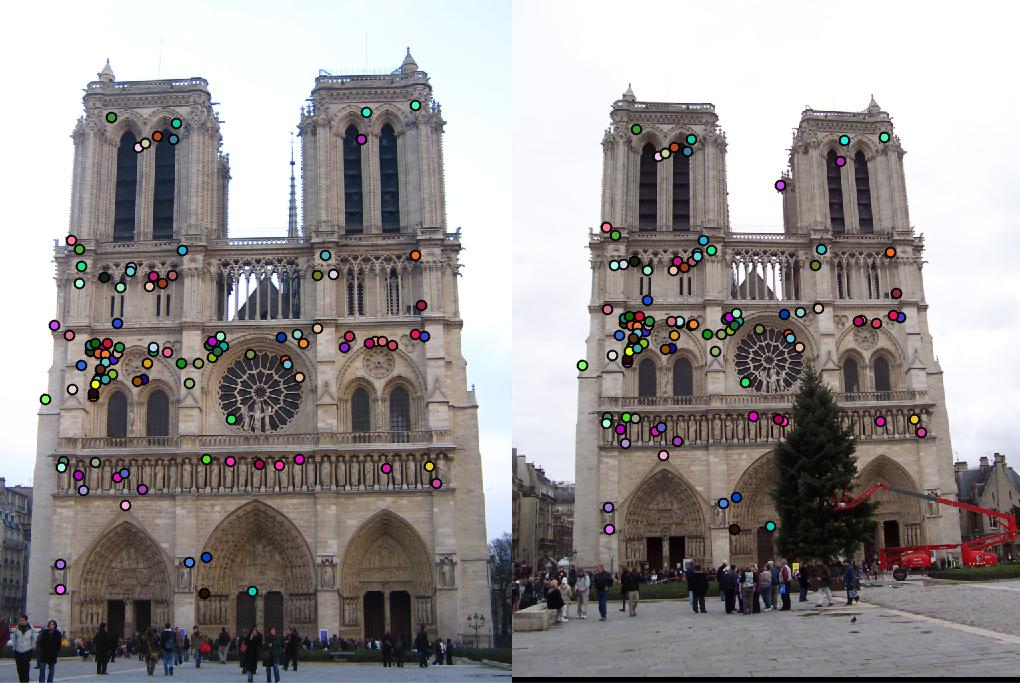 |
detected features of Notre Dame with threshold = 0.0001 and alpha = 0.04 |
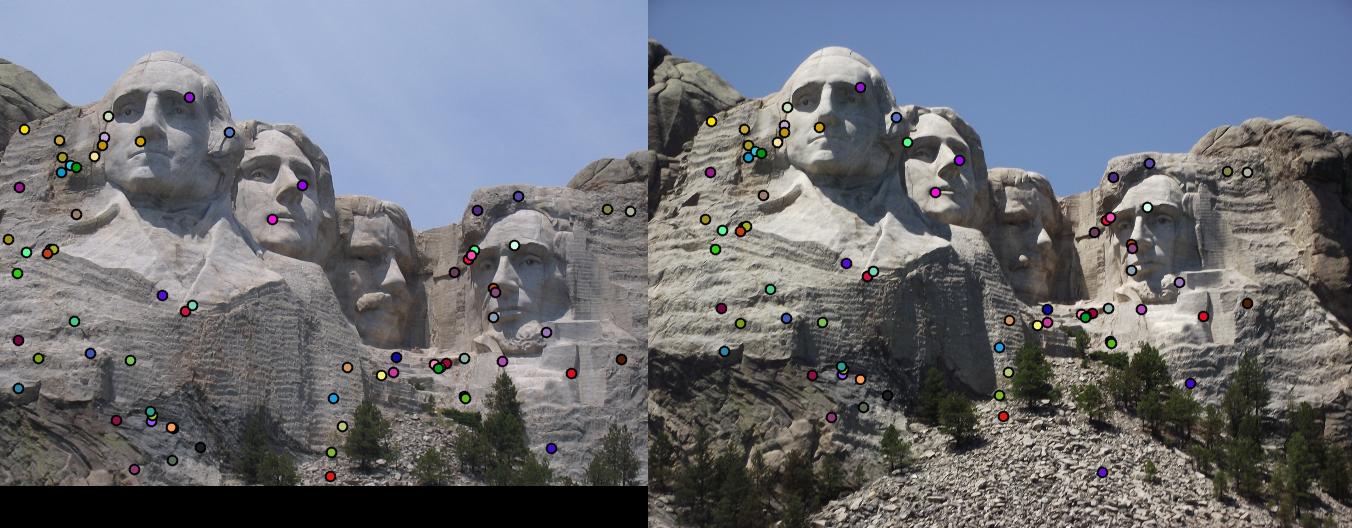 |
detected features of Mount Rushmore with threshold = 0.0001 and alpha = 0.04 |
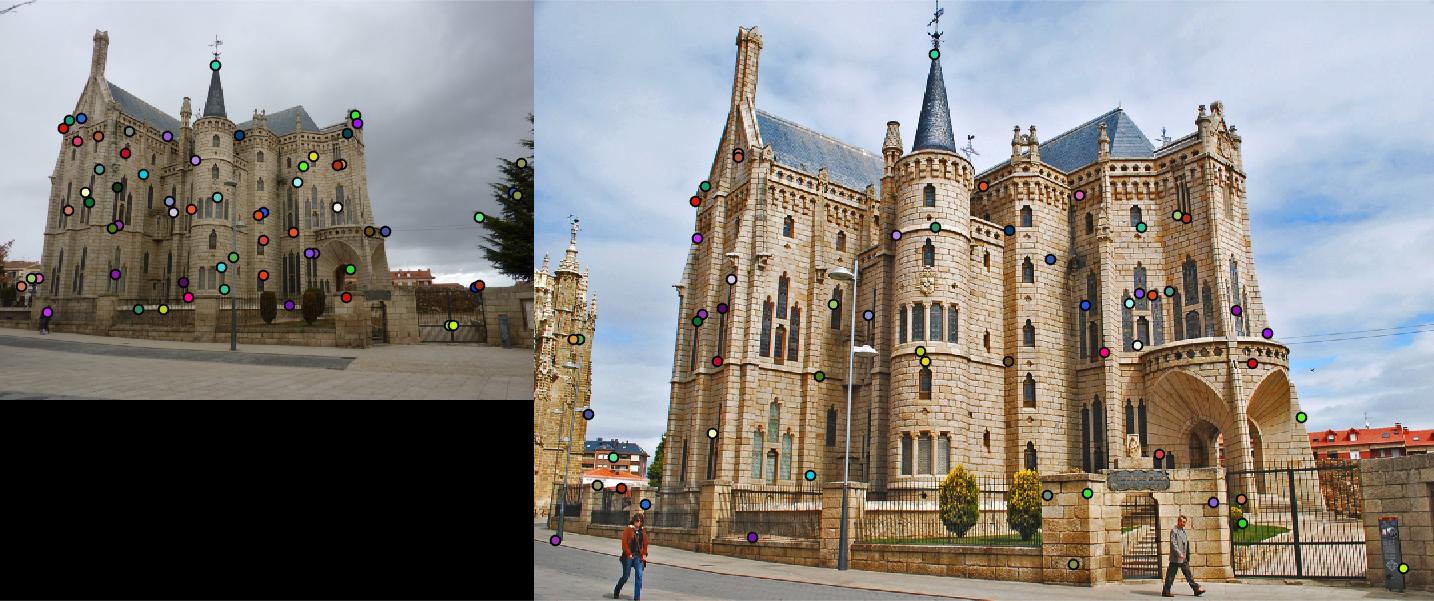 |
detected features of Episcopal Gaudi with threshold = 0.0001 and alpha = 0.06 |
3.Features matching
The thirs step is the matching of two images. For each descriptor in feature 1, calculate the distance between it and every descriptor in feature 2. Then, store all the results and sort the final array. Apply the nearest neighbout distance ratio to select the preferable matching points. The ratio is defined by the difference between the smallest distance between two descriptors in each features divided by the second smallest distance between other two. Finally, a threshold is set for choosing fitted points, in this case, 0.85
threshold = 0.85;
confidences = [];
matches = [];
k = 1;
for i = 1 : size(features1, 1)
for j = 1: size(features2, 1)
currentDist(j, :) = sum((features1(i,:) - features2(j,:)).^2);
end
[currentDist, indexlist] = sort(currentDist);
ratio = currentDist(1)/currentDist(2);
if ratio < threshold
matches(k, 1) = i;
matches(k, 2) = indexlist(1);
confidences(k) = ratio * 100;
k = k+1;
end
end
The ratio is stored for evaluation of the resulting image matching.
 |
 |
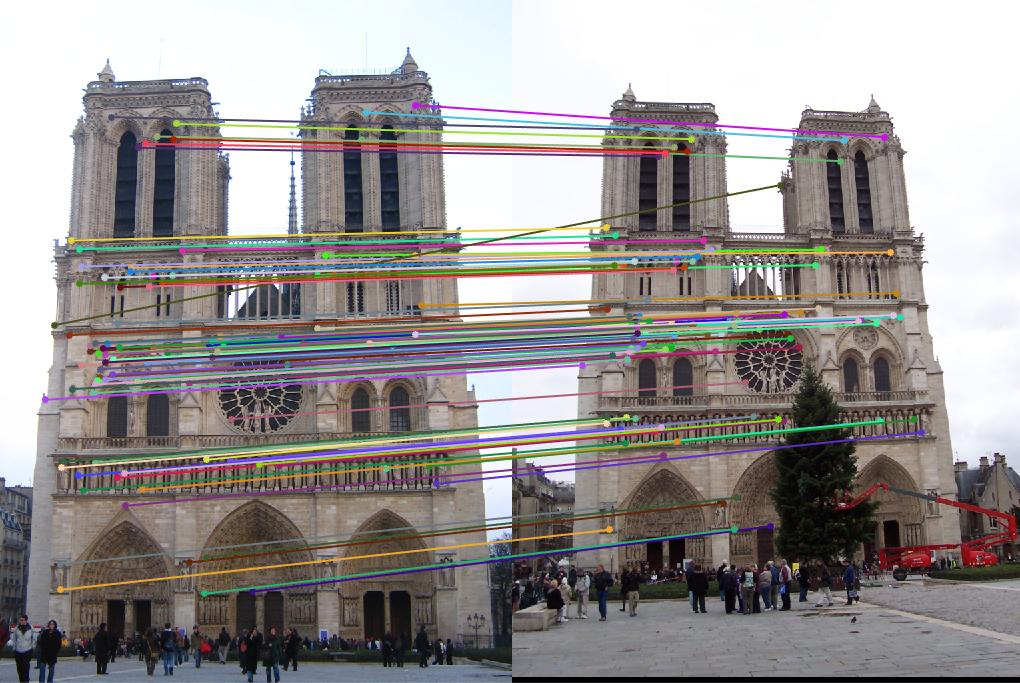
Notre Dame: threshold : 0.6 |
Result: 95 total good matches, 12 total bad matches. 89% accuracy |
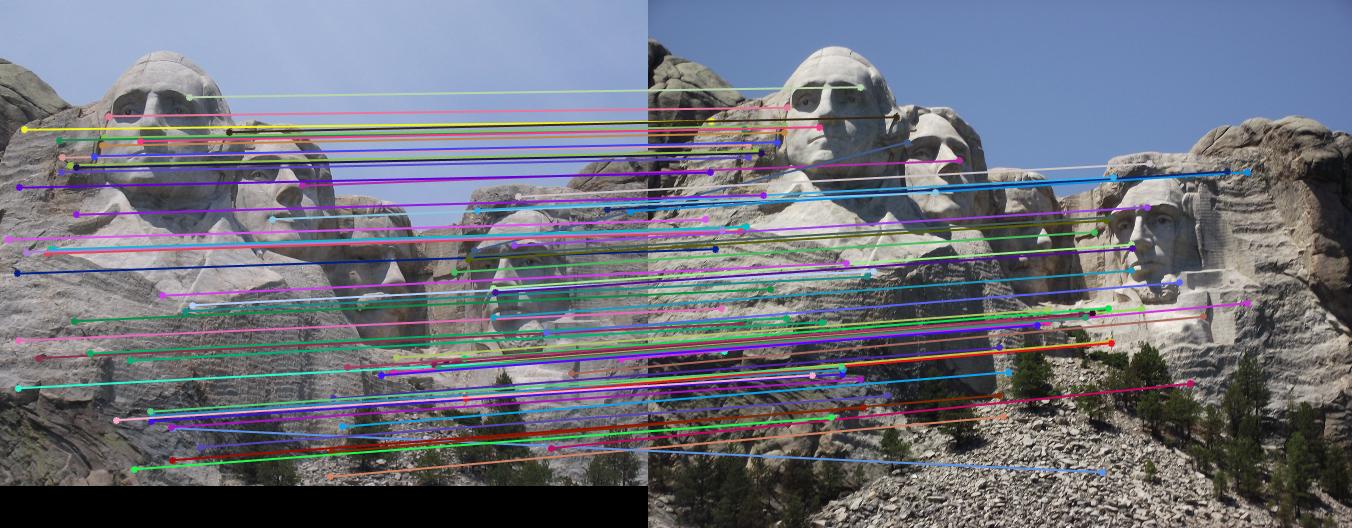 |
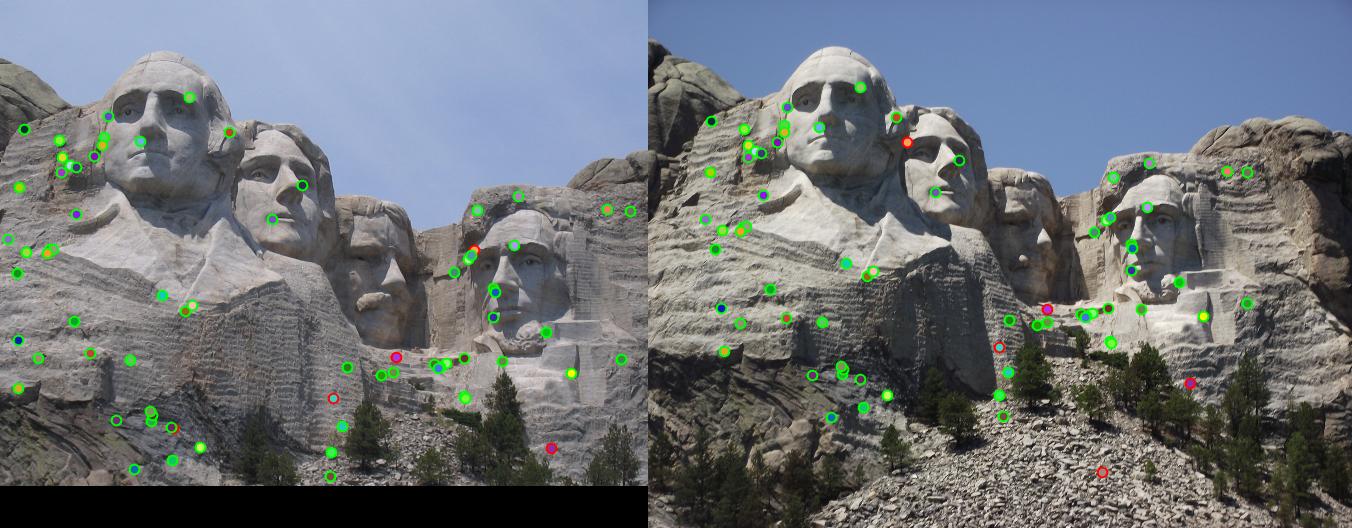 |
Mount Rushmore: threshold : 0.6 |
Result: 68 total good matches, 5 total bad matches. 93% accuracy |
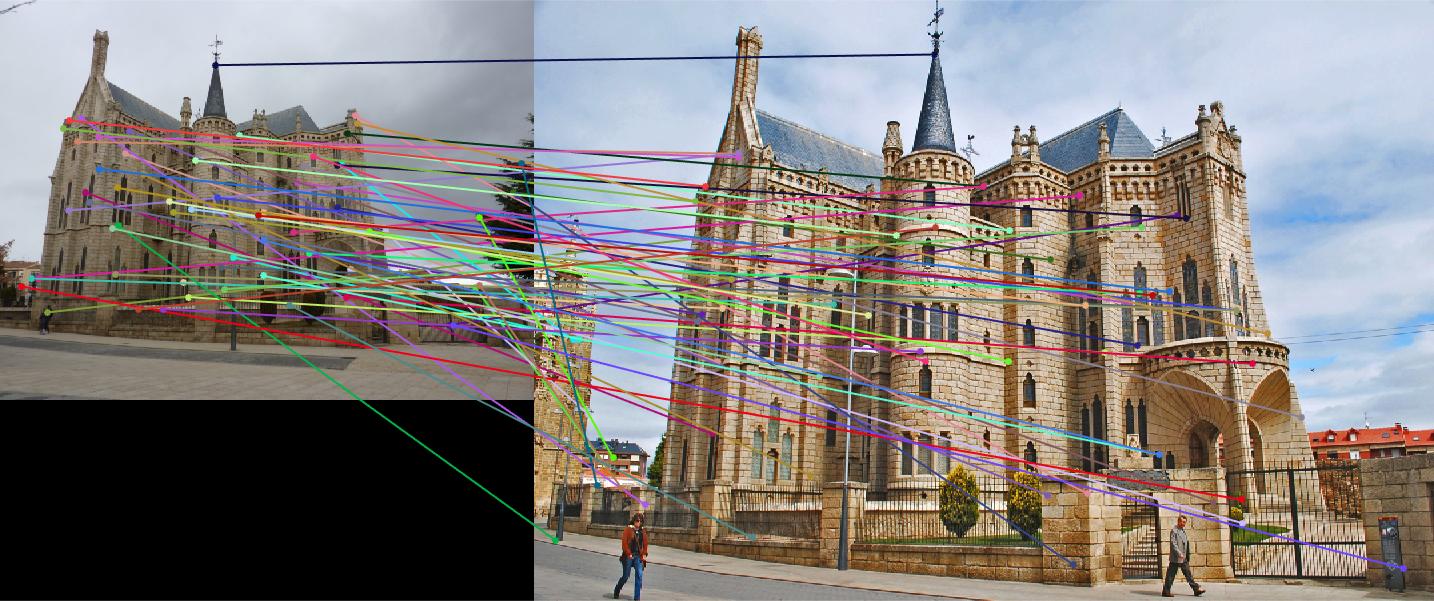 |
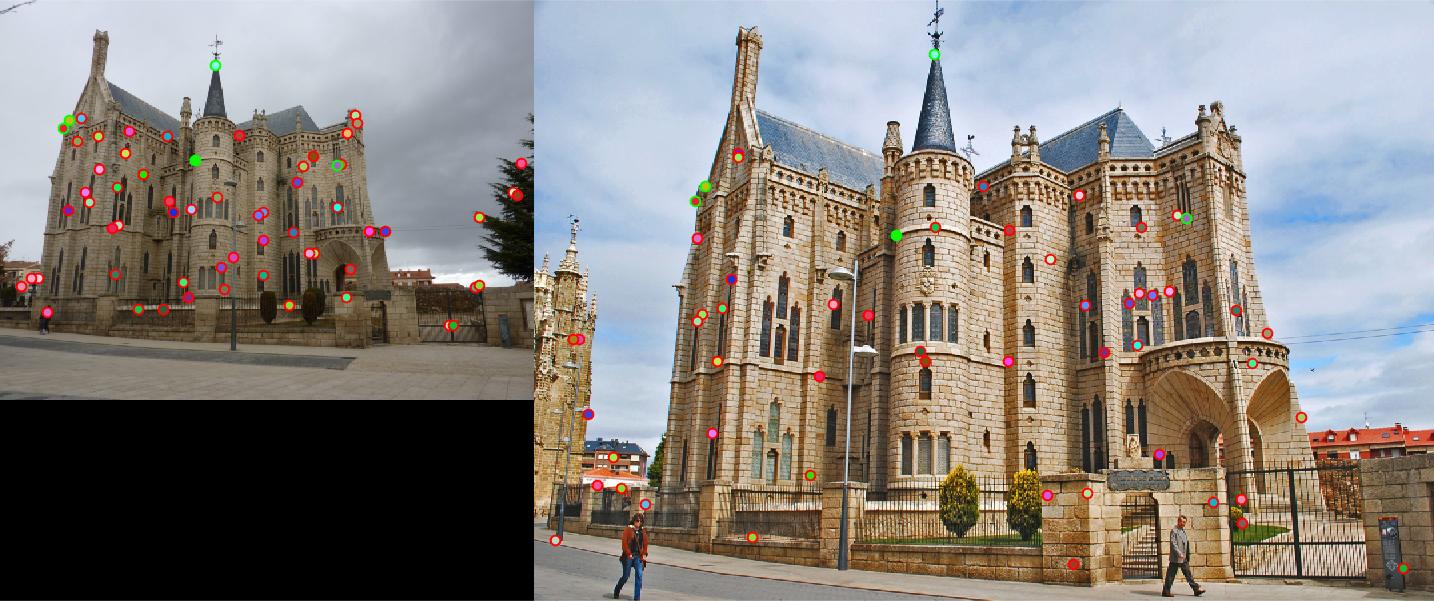 |
Episcopal Gaudi: threshold : 0.85 |
Result: 5 total good matches, 59 total bad matches. 8% accuracy |
4.Other examples
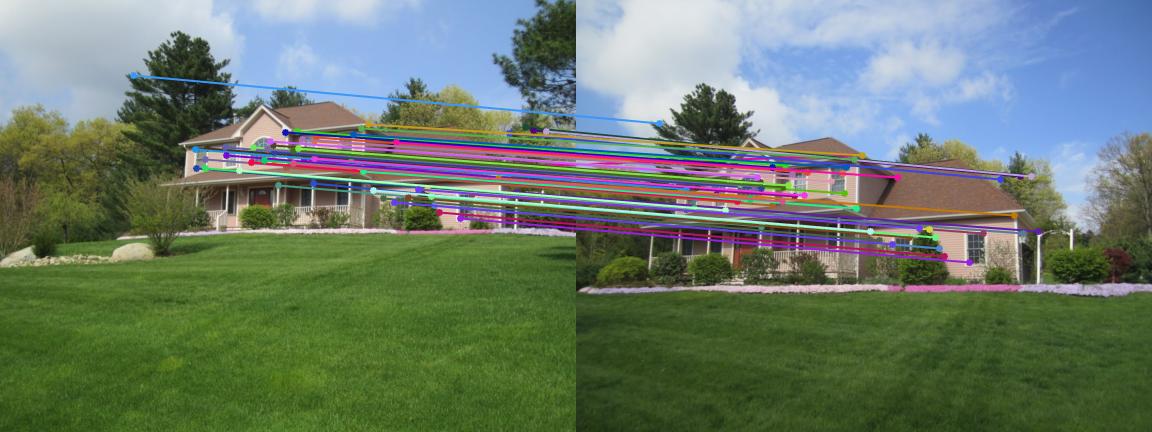 |
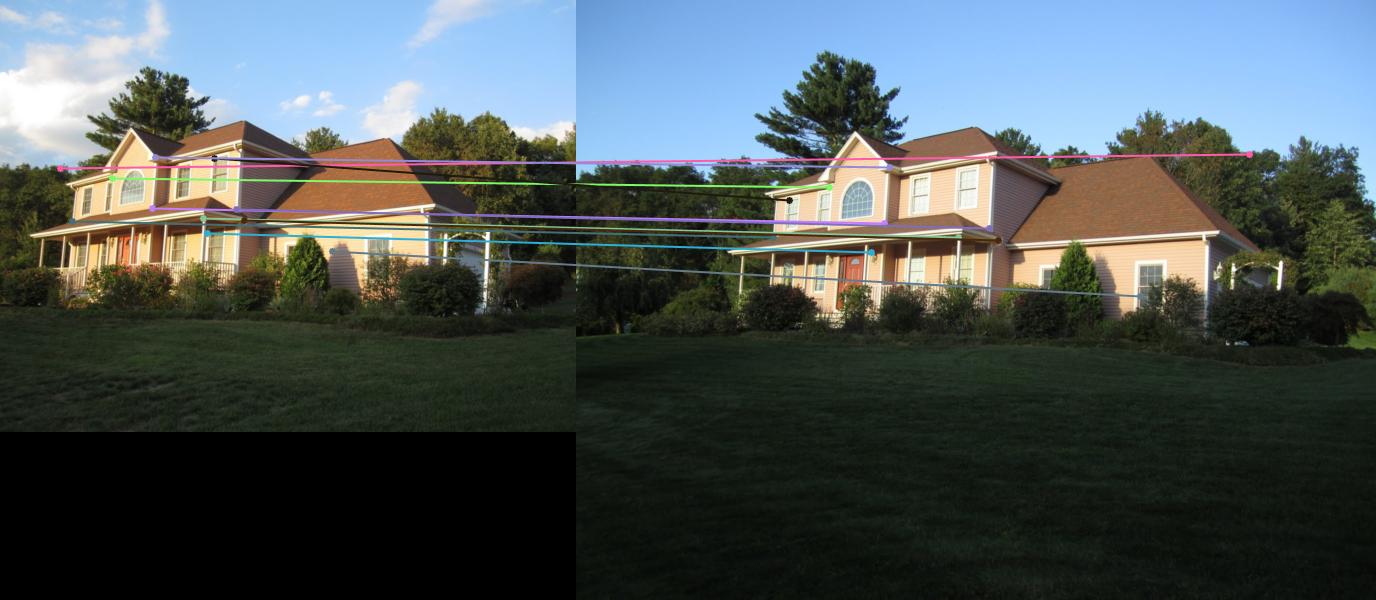 |
Example1 |
Example2 |
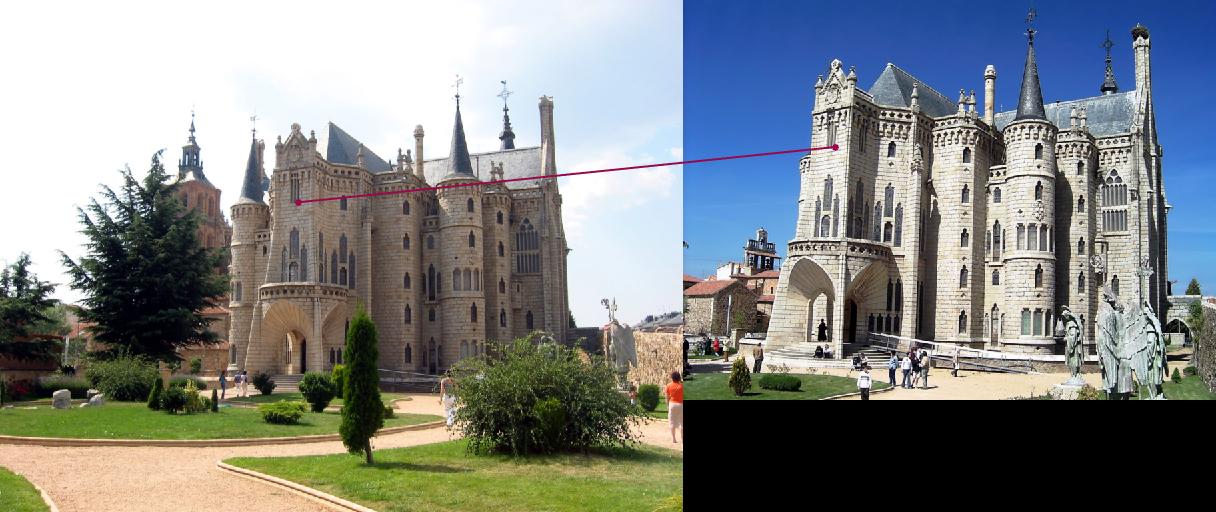 |
 |
Example3 |
Example4 |
5. Refection
The three provided images have the highest accuracy of Mount Rushmore, which is 93% and second highest of Notre Dame, which is 89%. For the Episcopal Gaudi, the accuracy is only 8%. After evaluating the resulting accuracy of the three provided images, it is found that images with similar size, in other word, resolution, similar brightness, orientation of object and environments would tend to achieve higher accuracy. The Eposcopal Gaudi attains poor accuracy because of sharp difference in brightness due to weather conditions and orientation of the construction. Using some of the examples in extra data, same pattern can be observed. In example 1, where the sunshine conditions and resolutions of the two images are nearly the same, there are large amount of matching dots. While in the example2 and example3, the conditions for sky are relatively different from each image, thus obtaining just a few lines. Difference in size of the images is also a reason for poor matching in example2 and example4.