Project 3 / Camera Calibration and Fundamental Matrix Estimation with RANSAC
This project had three parts:
- Camera Calibration
- Fundamental Matrix Estimation
- RANSAC and Fundamental Matrix
Camera Calibration
This required solving a simple system of linear equations. The first step was to build the system of equations. This was followed by solving using svd to find the appropraite matrix.
Example of code with highlighting
This is the MATLAB code I used to solve this problem.
x=zeros(size(Points_2D,1),4);
oneMat=ones(size(Points_2D,1),1);
Points_3D=[Points_3D,oneMat];
uMat=repmat(Points_2D(:,1),1,4);
vMat=repmat(Points_2D(:,2),1,4);
A=[Points_3D,x,-uMat.*Points_3D];
B=[x,Points_3D,-vMat.*Points_3D];
Mat=[A;B]
[U,S,V]=svd(Mat);
M=V(:,end);
M=reshape(M,[],3)';
Result of Part 1
The projection matrix is:| -0.4583, | 0.2947, | 0.0140, | -0.0040 |
| 0.0509, | 0.0546, | 0.5411, | 0.0524 |
| -0.1090, | -0.1783, | 0.0443, | -0.5968 |
The estimated location of camera is: -1.5127, -2.3517, 0.2826
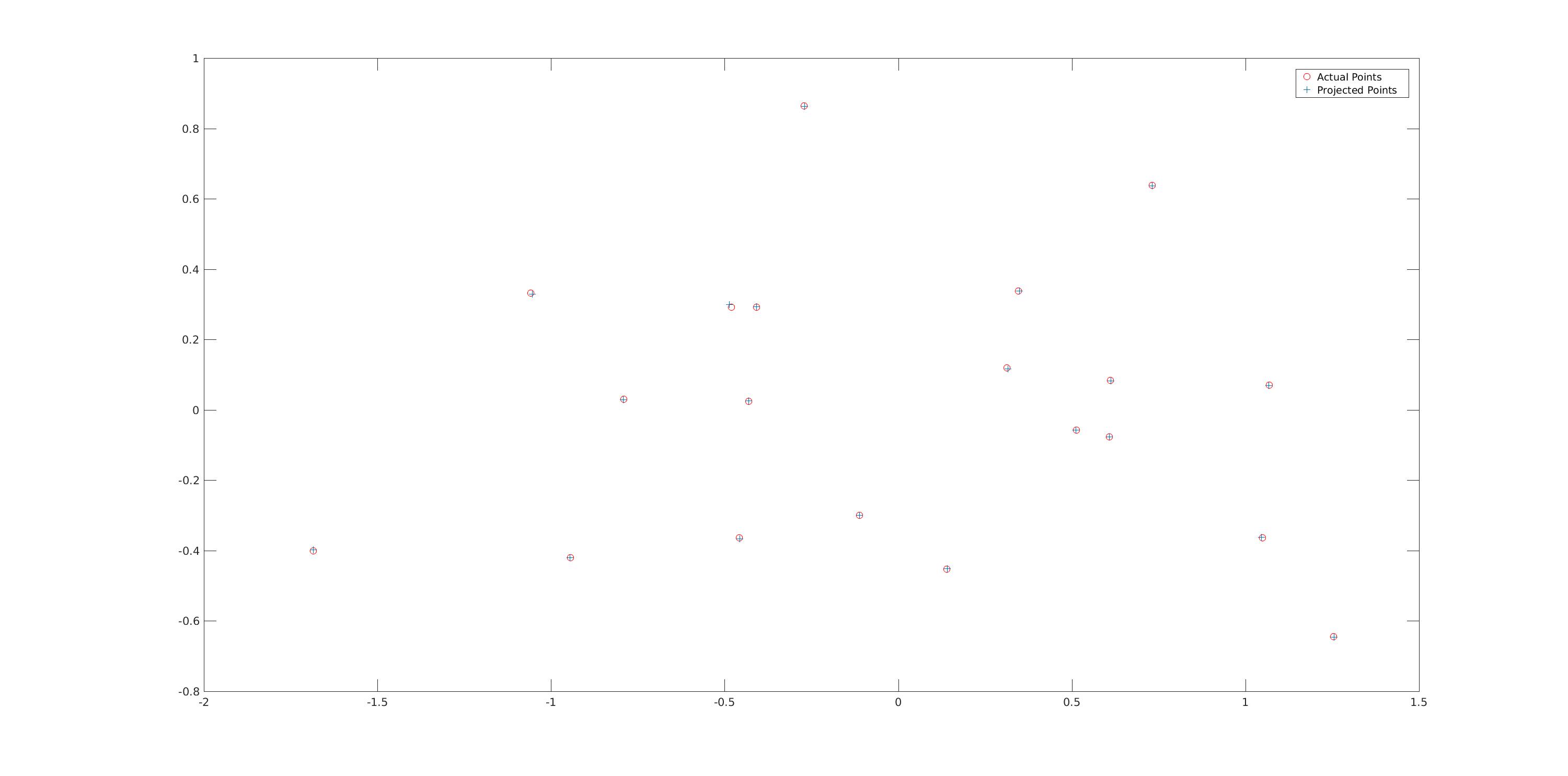
|
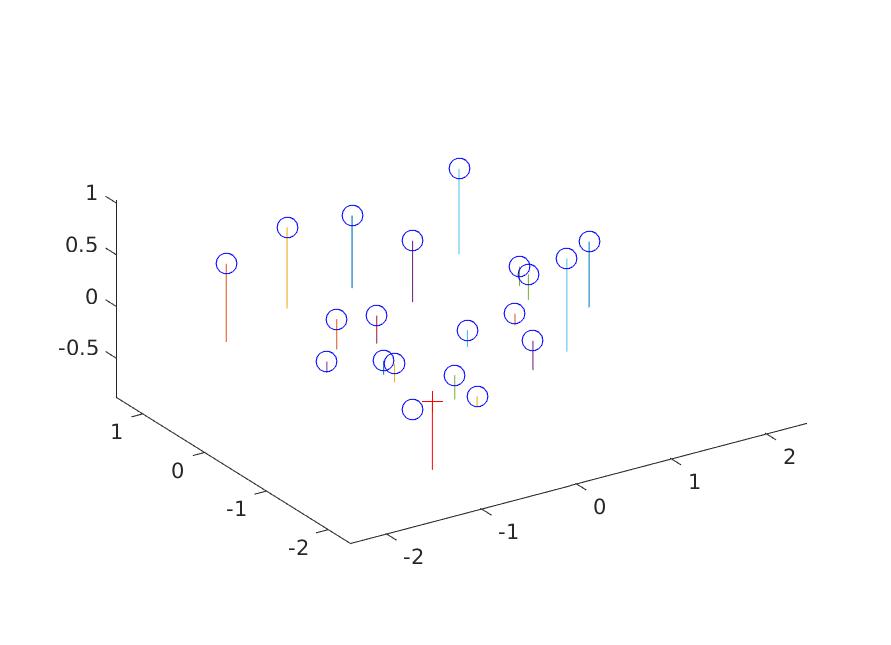
|
Fundamental Matrix Estimaton
This was again solving a set of linear equations and then using SVD
Result of Part 2
| -0.0000 | 0.0000 | -0.0019 |
| 0.0000 | 0.0000 | 0.0172 |
| -0.0009 | -0.0264 | 0.9995 |
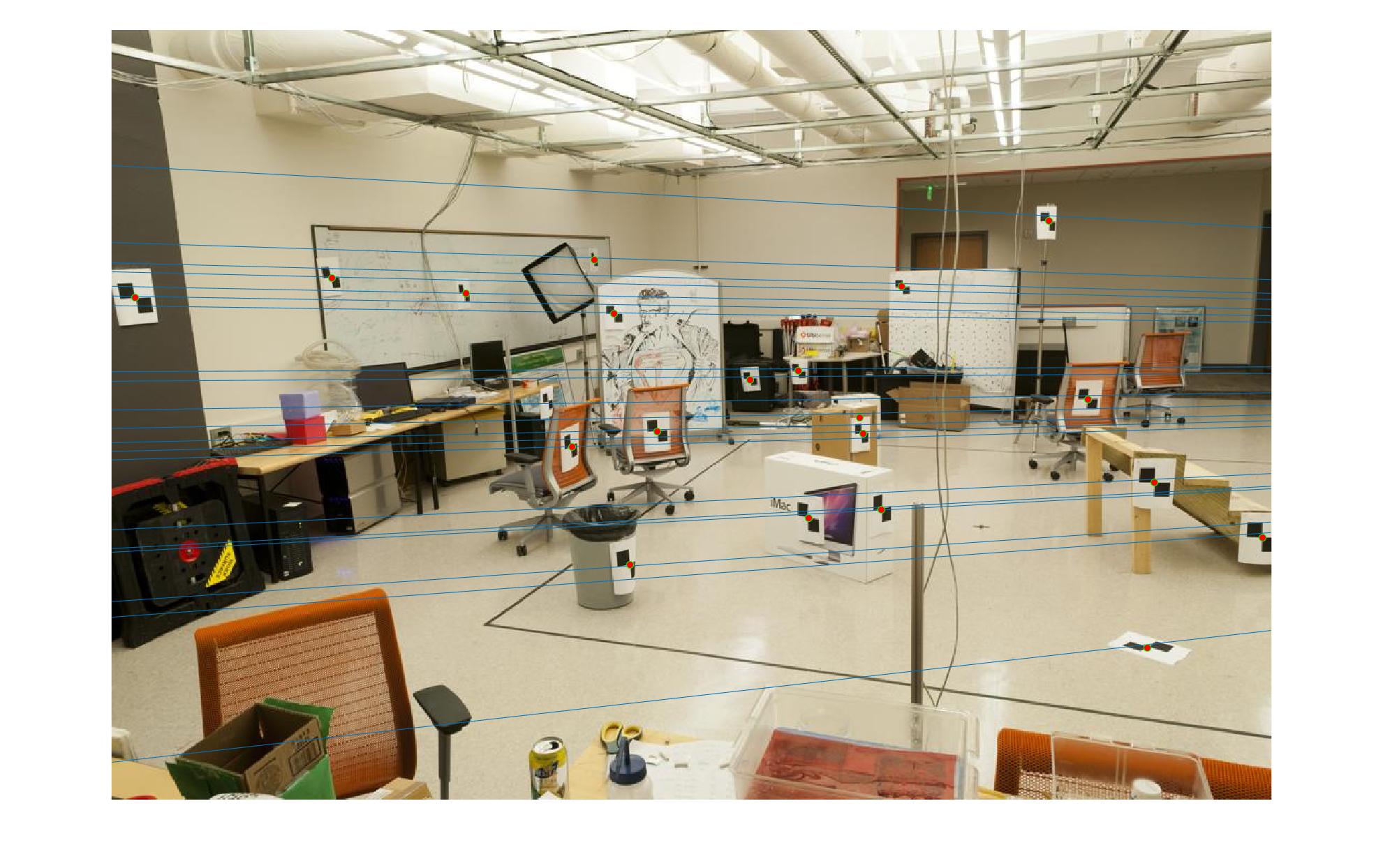

|
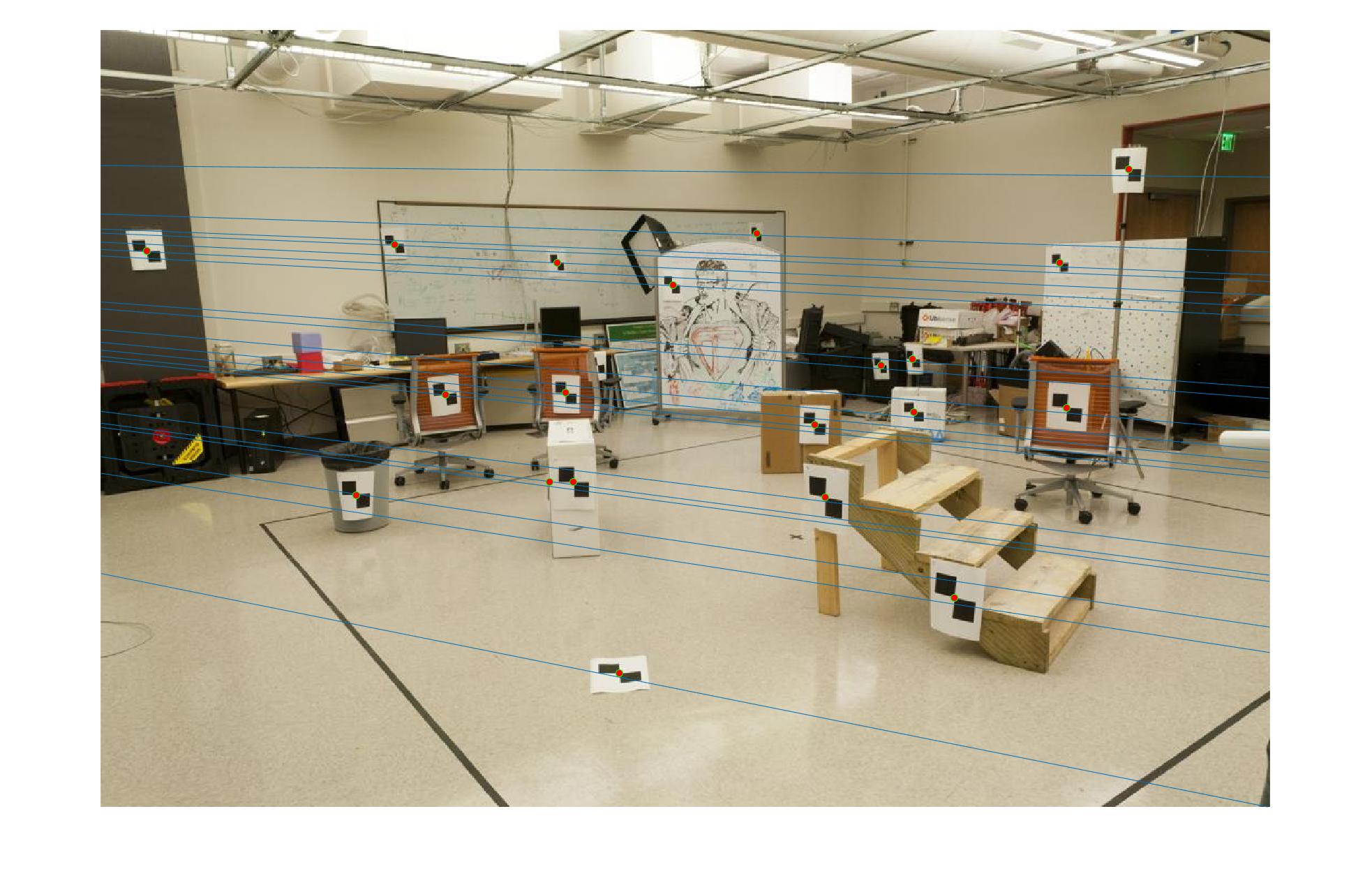

|
RANSAC and Fundamental Matrix Estimation
In order to accomplish this I ran 1000 iterations of RANSAC to find the best possible fundamental matrix by using 9 randomly chosed points to build a fundamental matrix and the finding the fundamental matrix with the maximum number of inliers.
I also used normlized points to get a better fundamental matrix and thus, better epipolar lines.
Results in a table
The first image in each image set is the image with non normalized points. The second image has normalized points.
Mount Rushmore

|
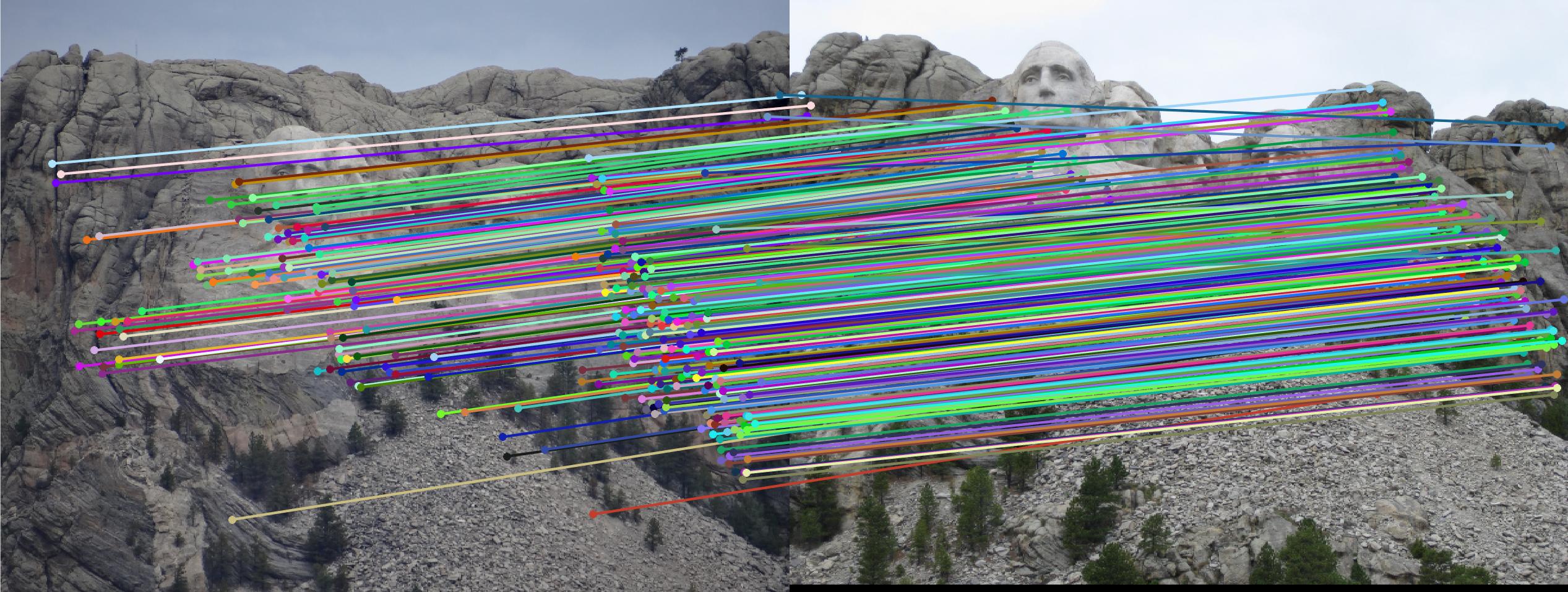

|
Notre Dame
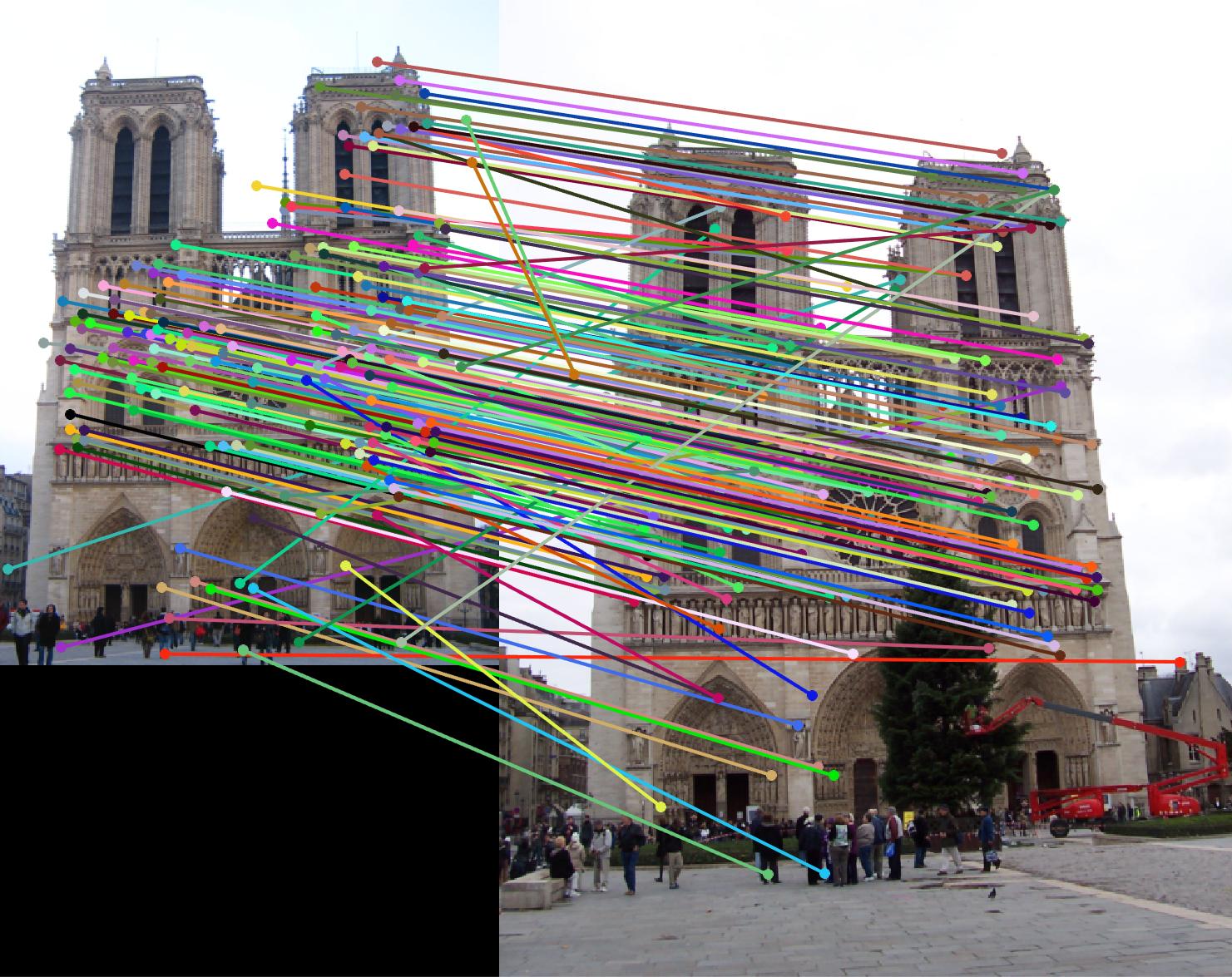

|
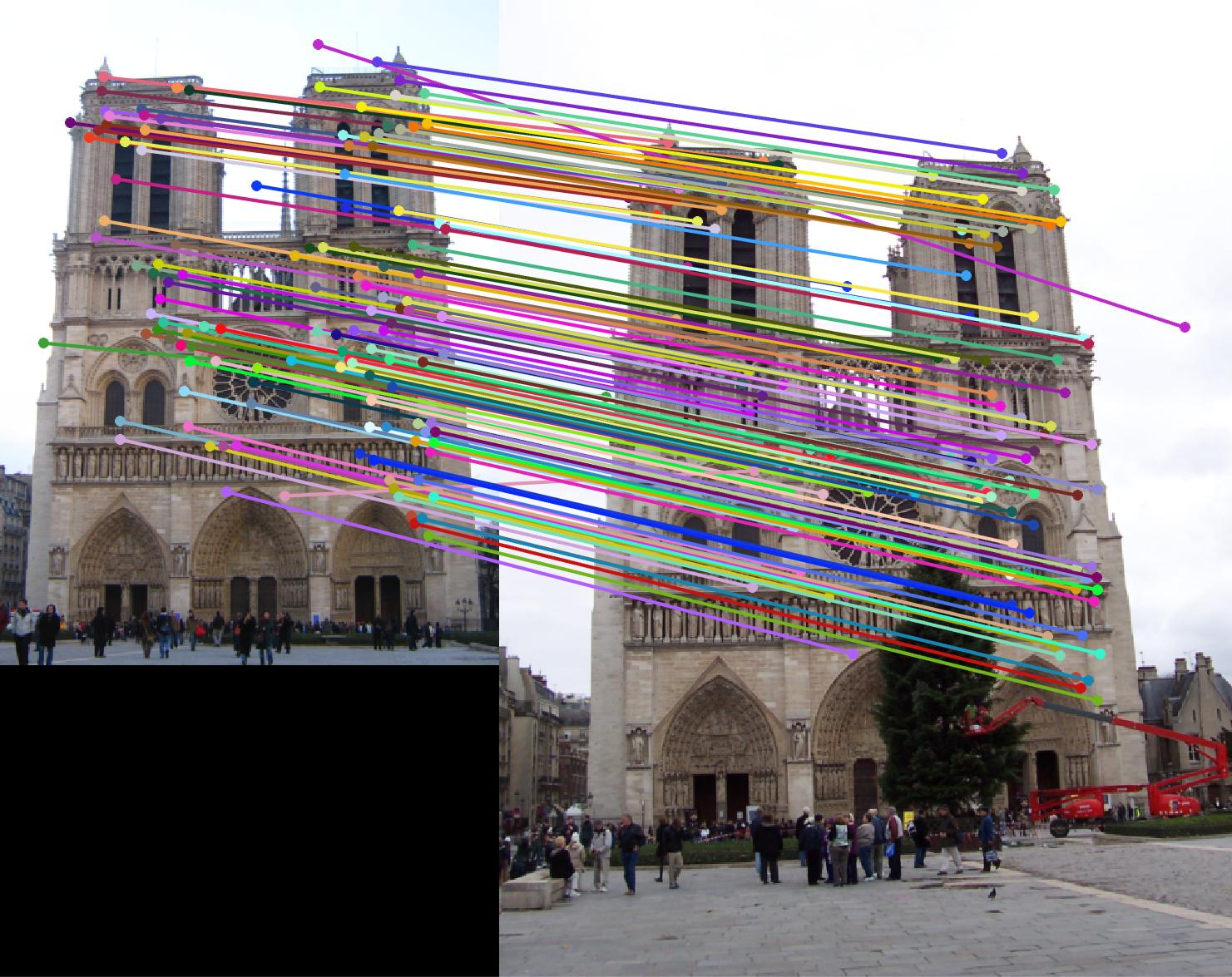

|
Episcopal Gaudi
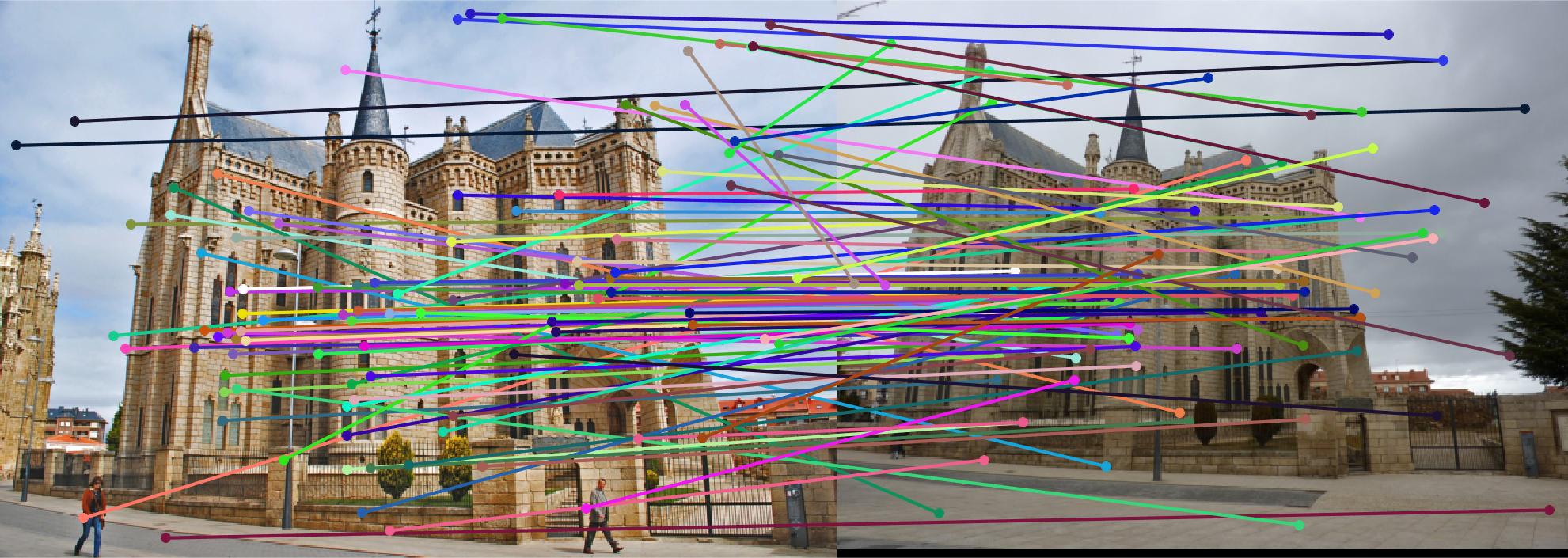

|
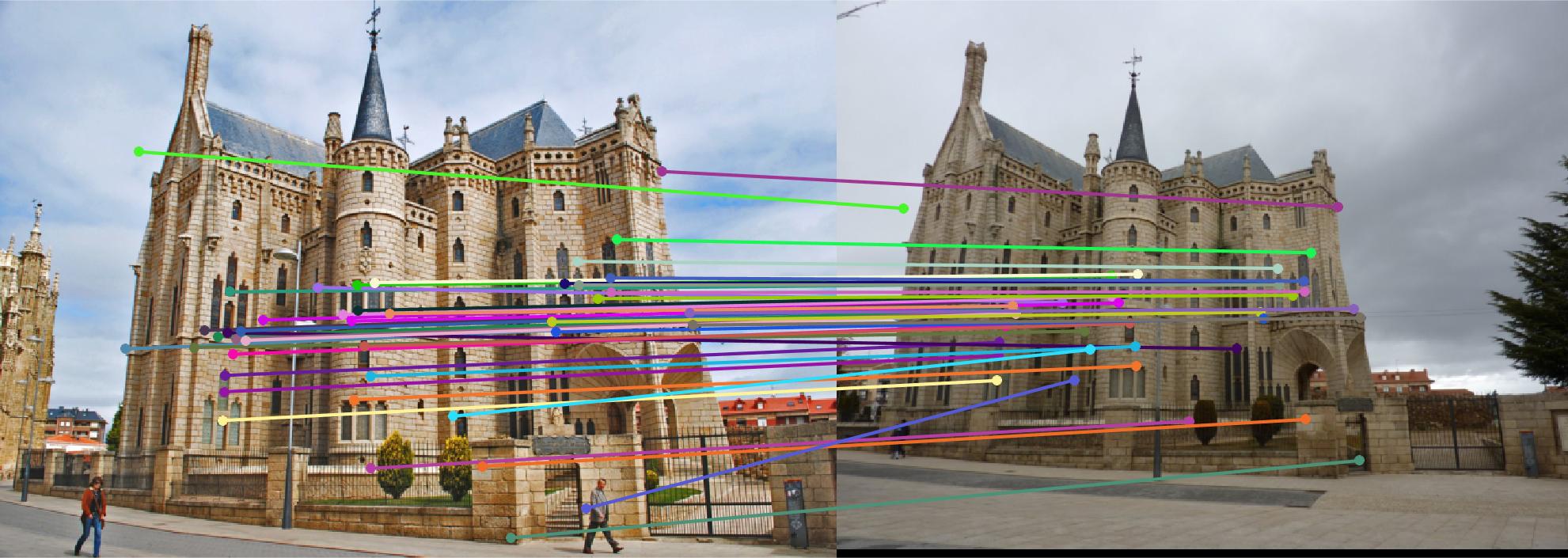

|
| Name | Features in image 1 | Features in image 2 | Matches | Inliers | Inliers with Normalization | threshold | |
|---|---|---|---|---|---|---|---|
| Mount Rushmore | 5583 | 5249 | 835 | 124 | 282 | 0.01 | |
| Notre Dame | 768 | 1633 | 266 | 266 | 180 | 0.1 | |
| Episcopal Gaudi | 1289 | 1035 | 126 | 96 | 42 | 0.1 |
Conclusions
Clear improvemnets can be seen with normalized points in all the image sets. Since random points are selected in every iteration to find out the fundamental matrix, the results in the final epipolar lines are not always the same. I have mentioned the best ones that I observed.
Also, varying threshold changes the number of inliers and consequnetly the best epipolar lines.