Project 3 / Camera Calibration and Fundamental Matrix Estimation with RANSAC
The aim of this project was to understand camera and scene geometry. Specifically, we estimateD the camera projection matrix, which maps 3D world coordinates to image coordinates, as well as the fundamental matrix, which relates points in one scene to epipolar lines in another.The three modules this project involved are
- Estimating Projection Matrix
- Estimating Fundamental Matrix
- Implementing RANSAC
- Extra credit: Normalizing points before running 9 point algorithm on them.
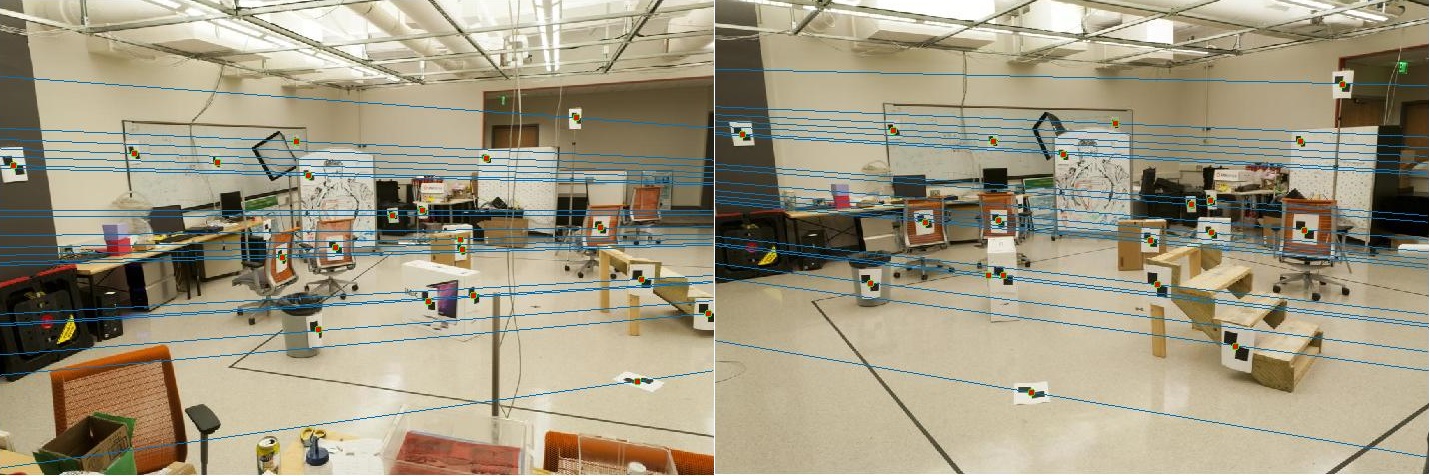

Epipolar lines for two images with different angles of view
Camera Calibration: Finding Projection Matrix
Projection matrix helps in mapping 3D world coordinates to 2D image coordinates. From the given set of 2D and 3D points, a homogeneous set of equations is set up. Projection matrix is a 3x4 matrix and to solve it, least squares method is used. The '\' operator of matlab has been made use of to do the same.
The resultant projection matrix that was computed is,
-0.7679 0.4938 0.0234 -0.0067
0.0852 0.0915 0.9065 0.0878
-0.1827 -0.2988 0.0742 -1.0000
Projection matrices can be scaled equivalents.
The above matrix can be obtained by multiplying the given normalized Projection matrix in project page by 1.675.
The residual obtained is 0.0445.
% To calculate the projection matrix - M
A = zeros(size(Points_3D,1)*2,11);
UV = zeros(size(Points_3D,1)*2,1);
for i=1:size(Points_3D,1)
X = Points_3D(i,1);
Y = Points_3D(i,2);
Z = Points_3D(i,3);
u = Points_2D(i,1);
v = Points_2D(i,2);
UV(i*2-1,1) = u;
UV(i*2,1) = v;
A(i*2-1,:)= [X, Y, Z, 1, 0, 0, 0, 0, -u*X, -u*Y, -u*Z];
A(i*2,:) = [0, 0, 0, 0, X, Y, Z, 1, -v*X, -v*Y, -v*Z];
end
M = A\UV;
M = M(:,1);
M = [M;1];
M = reshape(M,[],3)';
M = -1*M;
Using the projection matrix other intrinsic and extrinsic properties can be calculated. The centre of the camera location in world coordinates can be calculated by find the negation of product of last column of projection matrix and the inverse matrix of the first 3 columns of the projection matrix. The camera center that was computed using the above projection matrix is (-1.5126, -2.3517, 0.2827). This is almost equal to the camera center given in the project page.
%To calculate the camera center
Center = -1*inv(ProjectionMatrix(:,1:3))*ProjectionMatrix(:,end);
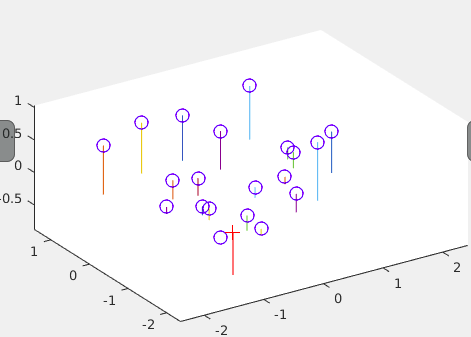
A visualisation of the camera center in the 3D scene.
Fundamental Matrix Estimation
Fundamental Matrix helps to estimate the mapping of points in one scene to the epipolar lines in the second scene. Given the corresponding 2D coordinate points in two images, the fundamental matrix can be calculated as
Fundamental Matrix Equation
Results for estimation without normalization are
The original transformation matrix is computed as the matrix product of transpose of transformation matrix of second image and normalized fundamental matrix and the transformation matrix of first image. The obtained Fundamental matrix:-0.0000 0.0000 -0.0019 0.0000 0.0000 0.0172 -0.0009 -0.0264 0.9995Epipolar lines:
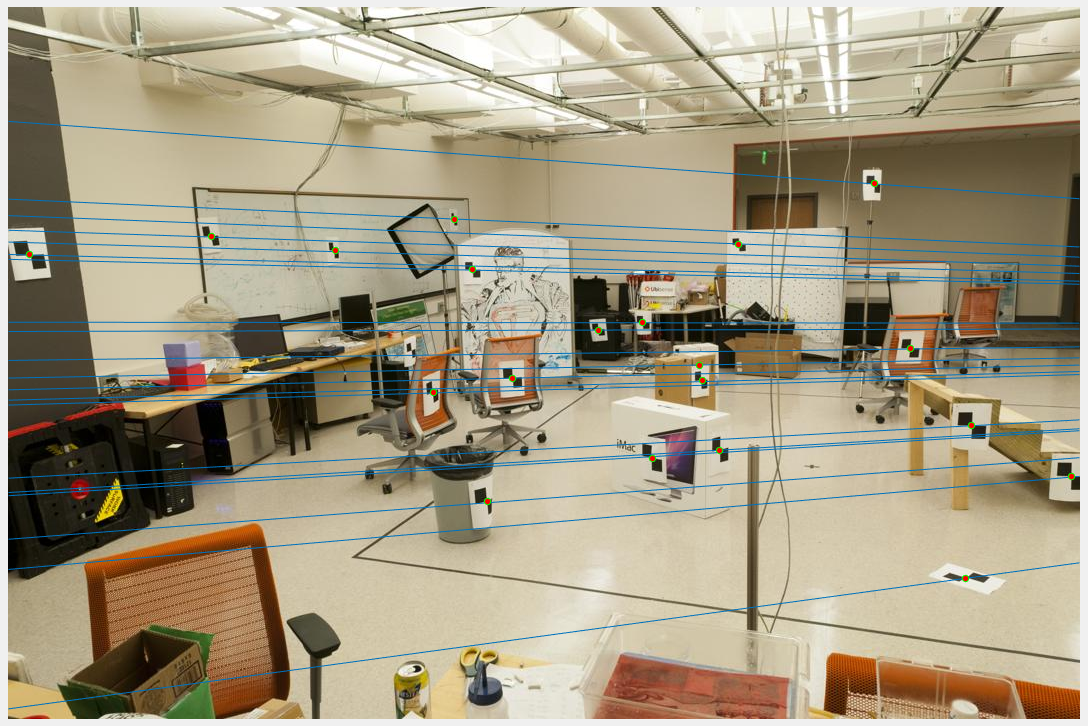
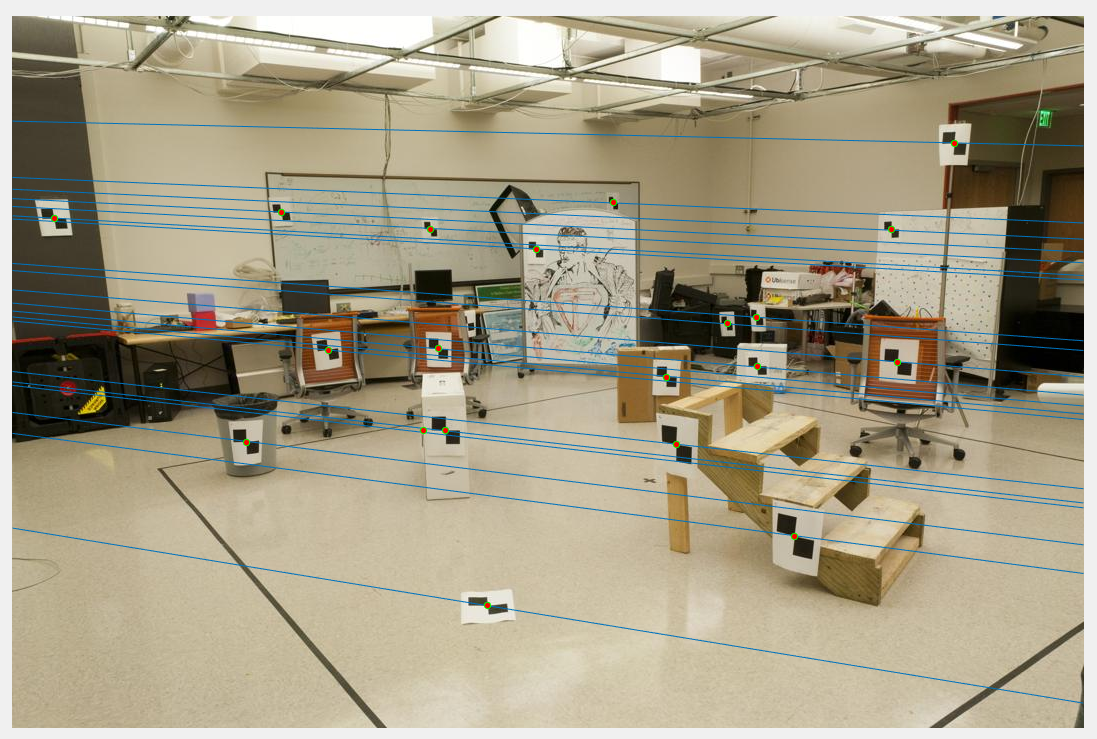


After normalisation:
The obtained fundamental matrix:-0.0000 0.0000 -0.0005 0.0000 -0.0000 0.0037 -0.0000 -0.0051 0.1192Epipolar lines:
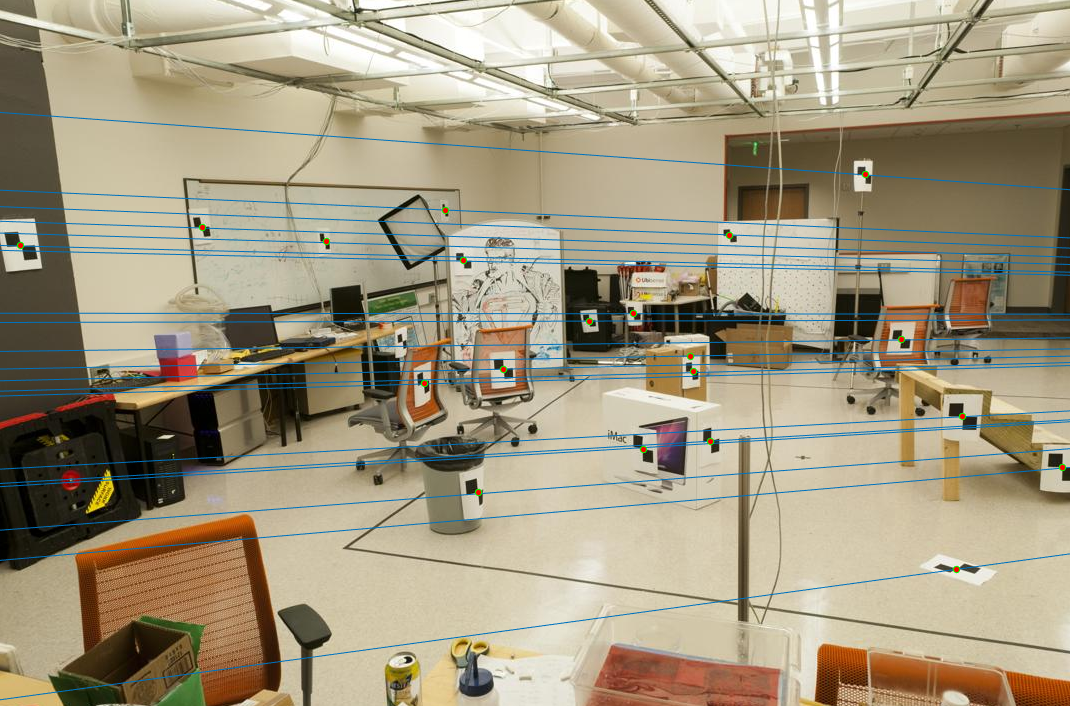
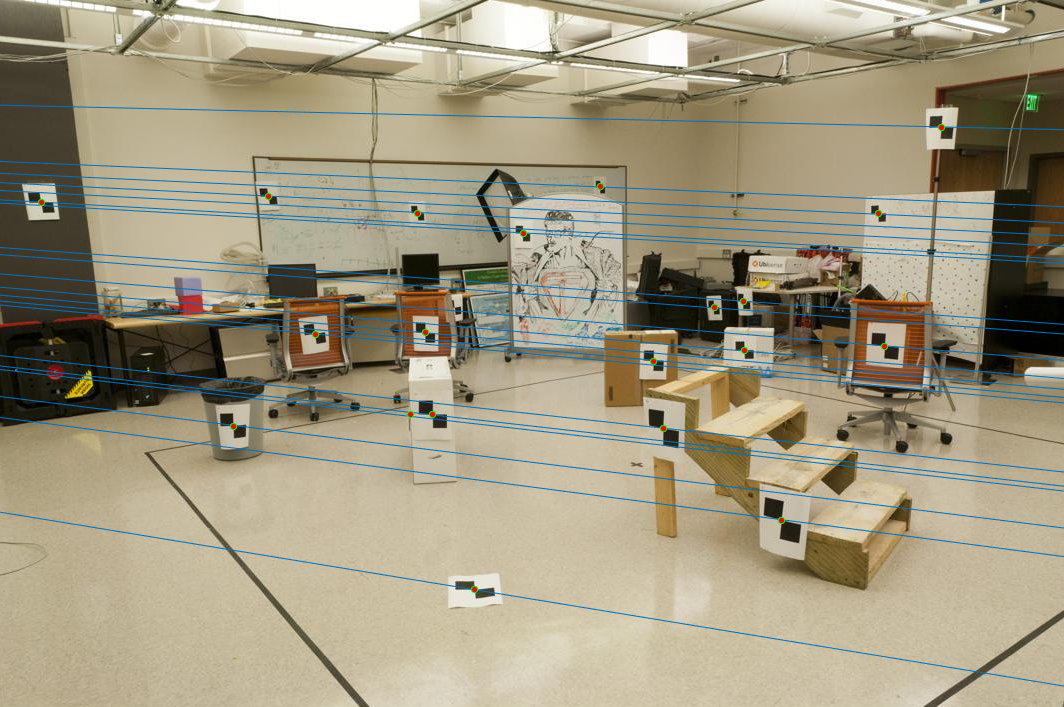


Estimating Epipolar Lines using RANSAC and Fundamental Matrix
Given two images, corresponding points in the two images and its fundamental matrix is needed to find the epipolar lines. SIFT is used to find the interest points in the image. The interest points might contain noisy data too. So they are not directly useful in find the fundamental matrix. To find the best fit model i.e. the best fundamental matrix for the found set of interest points, RANSAC is implemented. RANSAC picks the minimum number of points randomly inorder to estimate the model. Here 9 random points are picked without replacement to estimate the fundamental matrix. The method mentioned in the previous section is used to compute the fundamental matrix.
Once the fundamental matrix is calcuated, it is checked with all other interest points using the fundamental matrix equation mentioned in the previous section. Any point that satisfies the equation is known as an inlier. A threshold of 0.05 has been picked by trial and error method to differentiate between inliers and outliers. RANSAC is performed for 3000 iterations and the fundamental matrix with the highest number of inliers is selected as the best fundamental matrix.
% Step 1 - Sample numOfSamples points
samplePoints = randsample(size(matches_a,1),numOfSamples);
for i=1:numOfSamples
aPoints(i,:) = matches_a(samplePoints(i),:);
bPoints(i,:) = matches_b(samplePoints(i),:);
end
% Step 2 - Solve for all the points
tempFmatrix = estimate_fundamental_matrix(aPoints,bPoints);
% Step 3 - Score all the points
tempInlierCount = 0;
tempInlierIndex = zeros(size(matches_a,1),1);
for pt = 1:size(matches_a,1)
%calculate the inlying distance
A = [matches_a(pt,:)'; 1];
B = [matches_b(pt,:),1];
inlyingDist = B*tempFmatrix*A;
%Check if a point is inlier based on threshold value and inlying distance
if abs(inlyingDist) < Threshold
tempInlierCount = tempInlierCount + 1;
tempInlierIndex(pt,1) = 1;
end
end
%select the new FundamentalMatrix if it has more inliers than the old one
if tempInlierCount > numOfInliers
numOfInliers = tempInlierCount;
inlierIndices = tempInlierIndex;
FundamentalMatrix = tempFmatrix;
end
Results
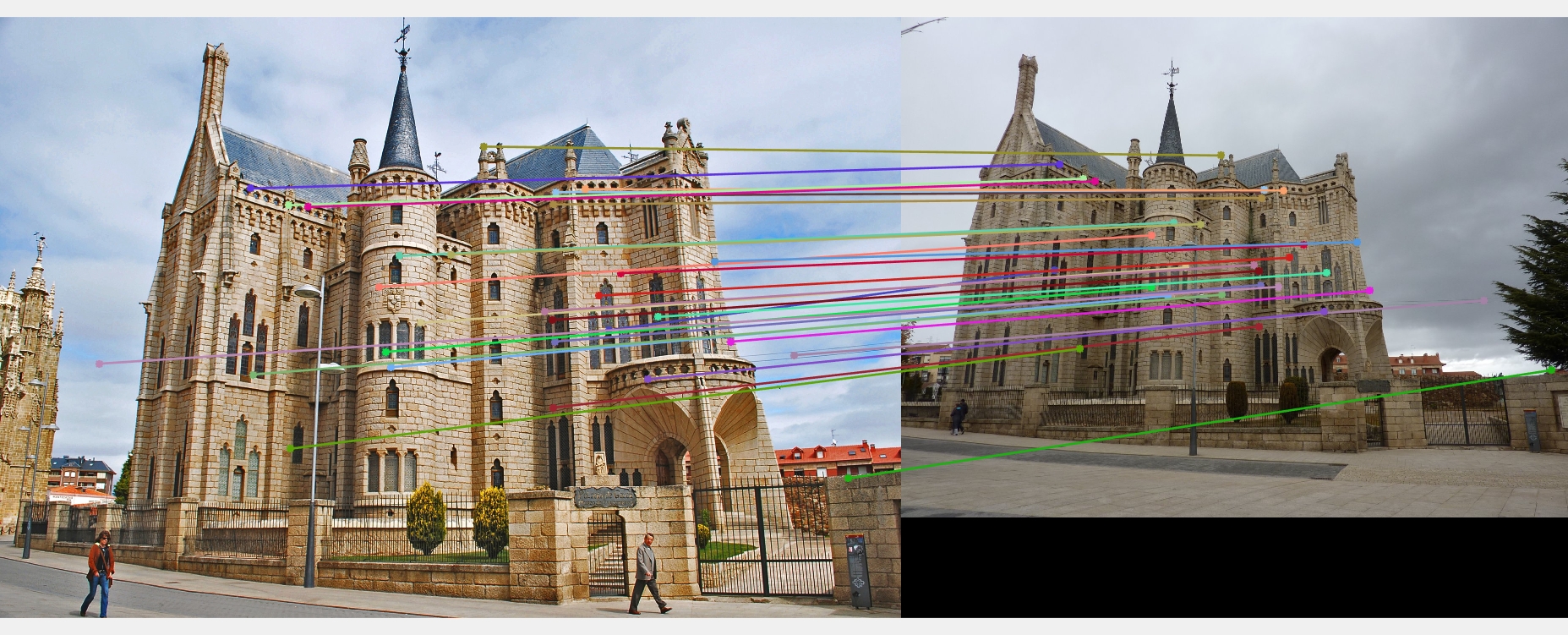
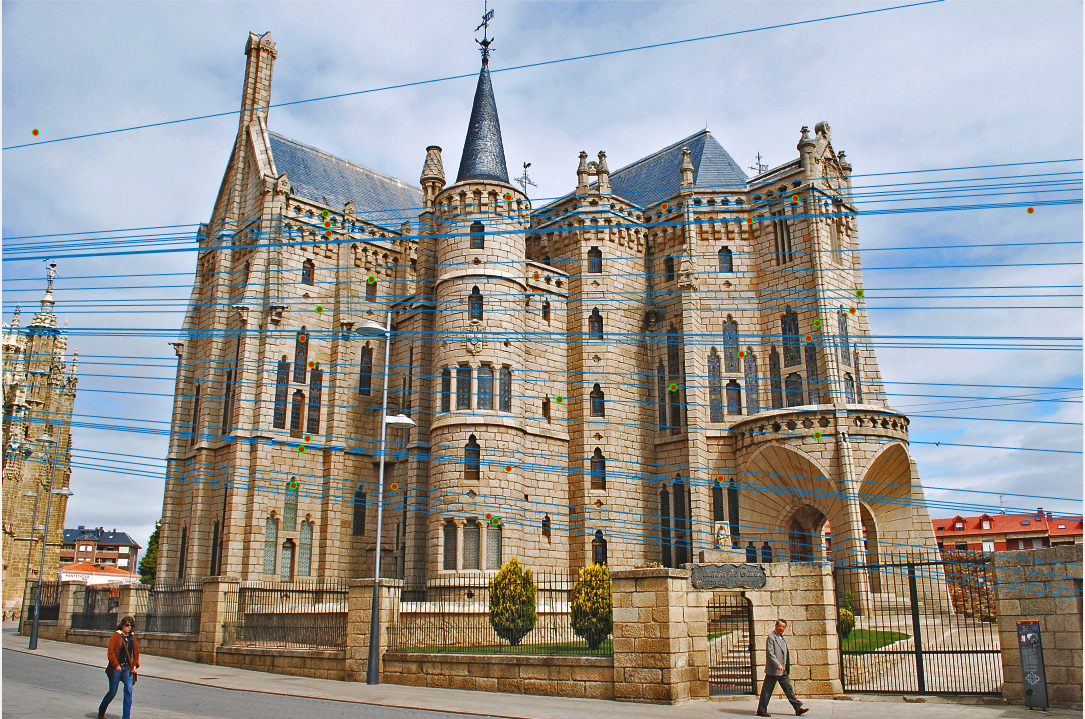
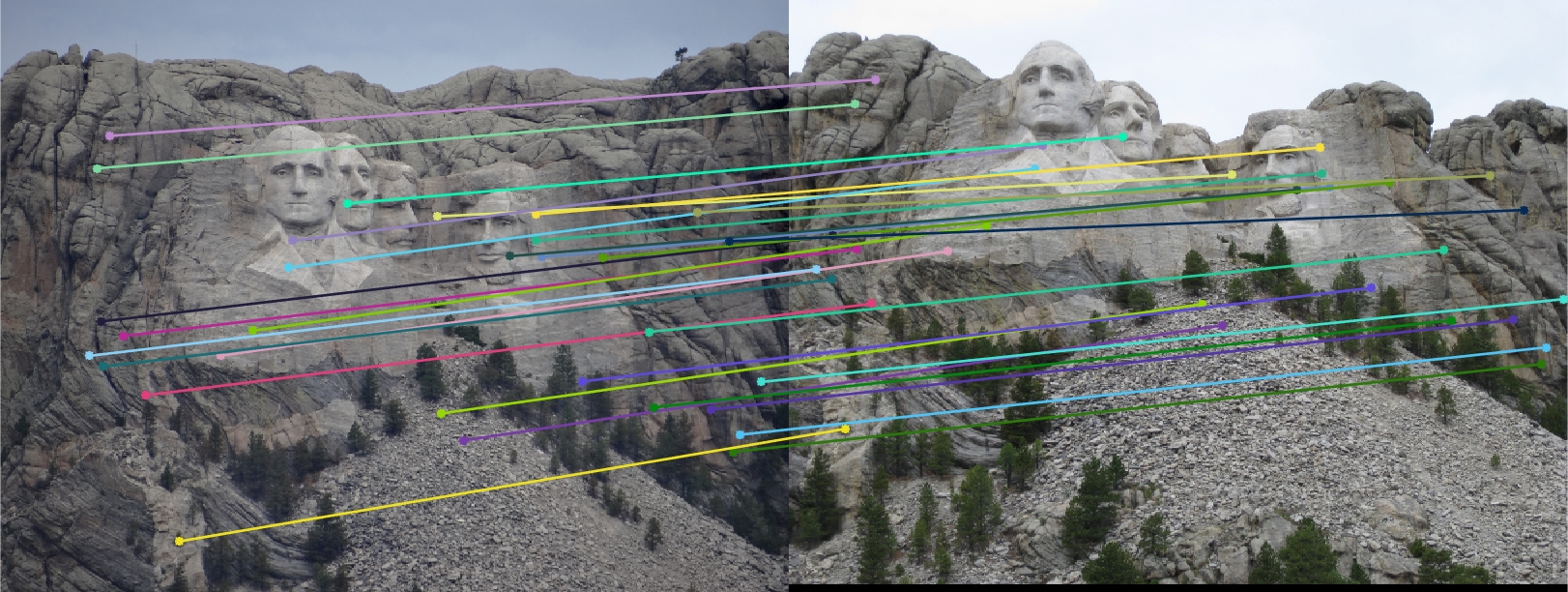
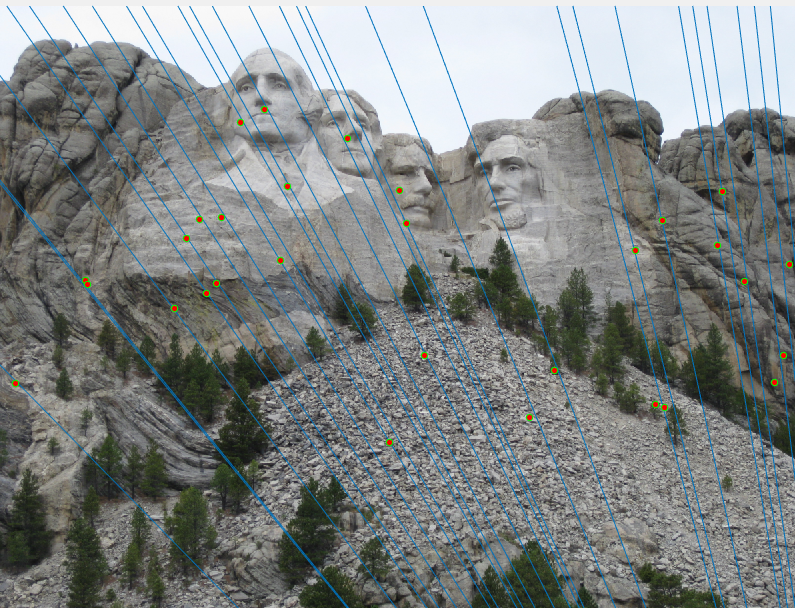
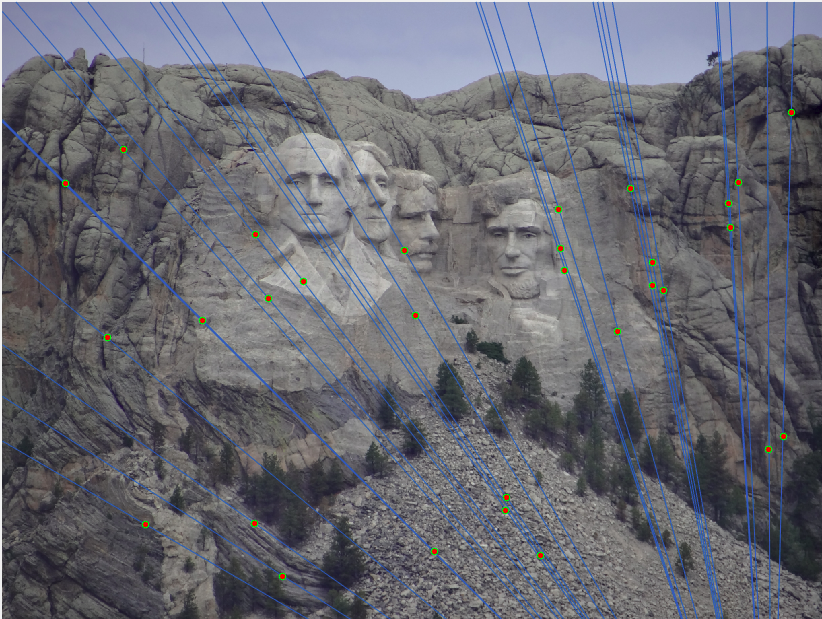
Only subset of points (30 points) are shown in the image for readability.Mount Rushmore (670 inliers)



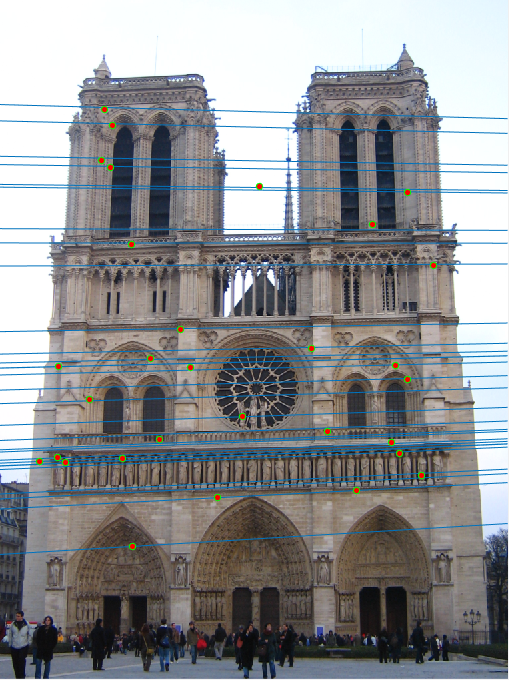
Notre Dame (688 inliers)
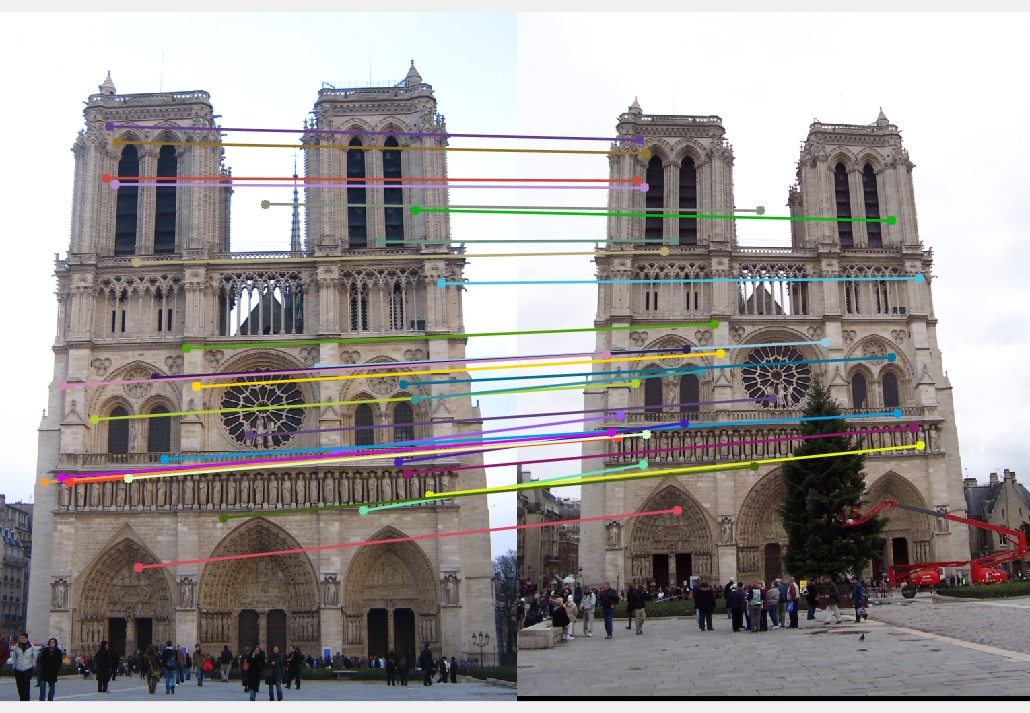
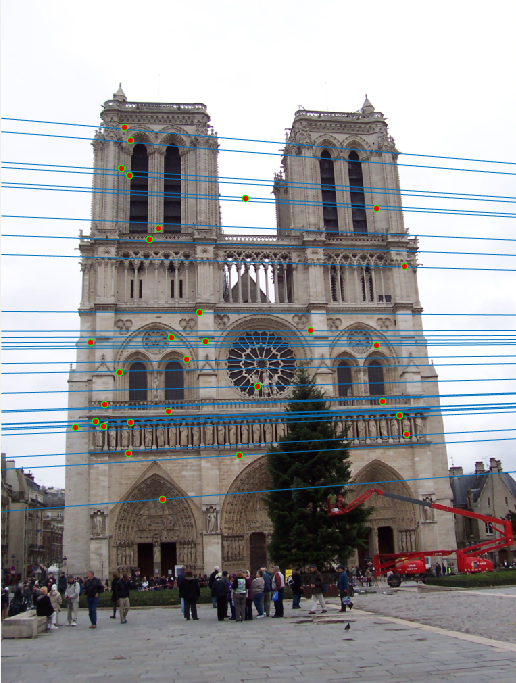



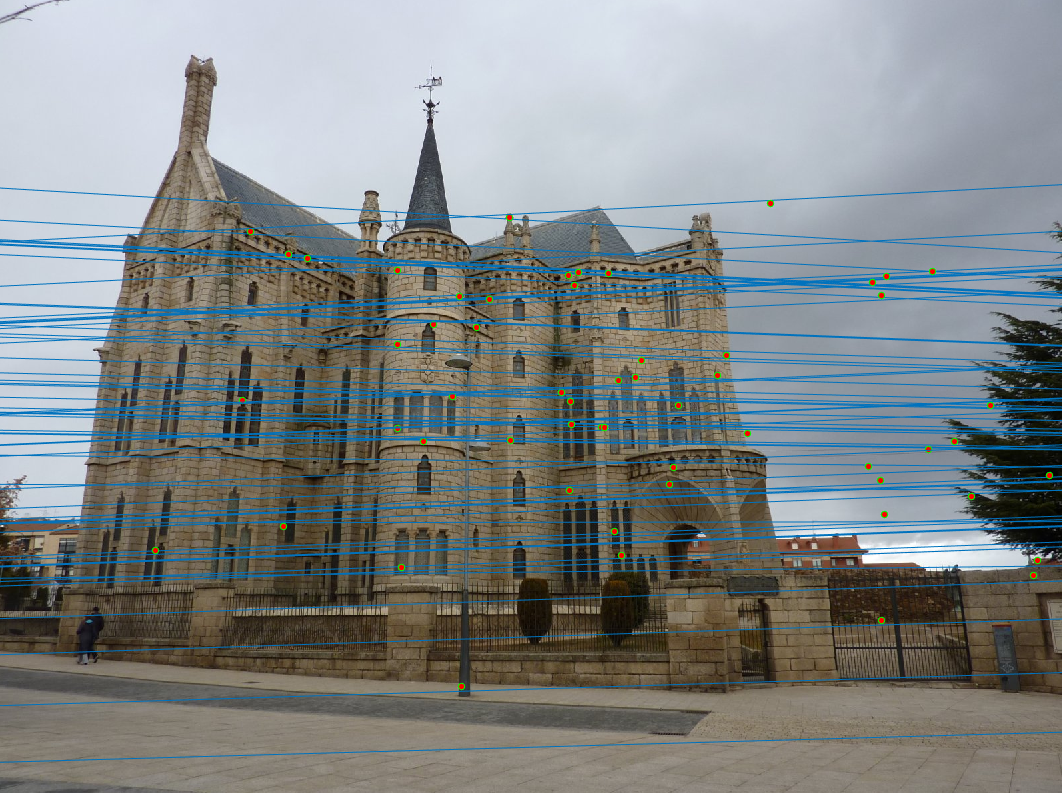
Episcopal Gaudi (539 inliers )


